Doc modernization (#3474)
* Change doc build to reST exclusively * Rewrite Intro doc in reST; create toctree * Update parameter and contribute * Convert tutorials to reST * Convert Python tutorials to reST * Convert CLI and Julia docs to reST * Enable markdown for R vignettes * Done migrating to reST * Add guzzle_sphinx_theme to requirements * Add breathe to requirements * Fix search bar * Add link to user forum
This commit is contained in:
committed by
 Philip Cho
Philip Cho
parent
d0f45bede0
commit
e19dded9a3
@@ -1,187 +0,0 @@
|
||||
Distributed XGBoost YARN on AWS
|
||||
===============================
|
||||
This is a step-by-step tutorial on how to setup and run distributed [XGBoost](https://github.com/dmlc/xgboost)
|
||||
on an AWS EC2 cluster. Distributed XGBoost runs on various platforms such as MPI, SGE and Hadoop YARN.
|
||||
In this tutorial, we use YARN as an example since this is a widely used solution for distributed computing.
|
||||
|
||||
Prerequisite
|
||||
------------
|
||||
We need to get a [AWS key-pair](http://docs.aws.amazon.com/AWSEC2/latest/UserGuide/ec2-key-pairs.html)
|
||||
to access the AWS services. Let us assume that we are using a key ```mykey``` and the corresponding permission file ```mypem.pem```.
|
||||
|
||||
We also need [AWS credentials](http://docs.aws.amazon.com/AWSSimpleQueueService/latest/SQSGettingStartedGuide/AWSCredentials.html),
|
||||
which includes an `ACCESS_KEY_ID` and a `SECRET_ACCESS_KEY`.
|
||||
|
||||
Finally, we will need a S3 bucket to host the data and the model, ```s3://mybucket/```
|
||||
|
||||
Setup a Hadoop YARN Cluster
|
||||
---------------------------
|
||||
This sections shows how to start a Hadoop YARN cluster from scratch.
|
||||
You can skip this step if you have already have one.
|
||||
We will be using [yarn-ec2](https://github.com/tqchen/yarn-ec2) to start the cluster.
|
||||
|
||||
We can first clone the yarn-ec2 script by the following command.
|
||||
```bash
|
||||
git clone https://github.com/tqchen/yarn-ec2
|
||||
```
|
||||
|
||||
To use the script, we must set the environment variables `AWS_ACCESS_KEY_ID` and
|
||||
`AWS_SECRET_ACCESS_KEY` properly. This can be done by adding the following two lines in
|
||||
`~/.bashrc` (replacing the strings with the correct ones)
|
||||
|
||||
```bash
|
||||
export AWS_ACCESS_KEY_ID=AKIAIOSFODNN7EXAMPLE
|
||||
export AWS_SECRET_ACCESS_KEY=wJalrXUtnFEMI/K7MDENG/bPxRfiCYEXAMPLEKEY
|
||||
```
|
||||
|
||||
Now we can launch a master machine of the cluster from EC2
|
||||
```bash
|
||||
./yarn-ec2 -k mykey -i mypem.pem launch xgboost
|
||||
```
|
||||
Wait a few mininutes till the master machine gets up.
|
||||
|
||||
After the master machine gets up, we can query the public DNS of the master machine using the following command.
|
||||
```bash
|
||||
./yarn-ec2 -k mykey -i mypem.pem get-master xgboost
|
||||
```
|
||||
It will show the public DNS of the master machine like ```ec2-xx-xx-xx.us-west-2.compute.amazonaws.com```
|
||||
Now we can open the browser, and type (replace the DNS with the master DNS)
|
||||
```
|
||||
ec2-xx-xx-xx.us-west-2.compute.amazonaws.com:8088
|
||||
```
|
||||
This will show the job tracker of the YARN cluster. Note that we may have to wait a few minutes before the master finishes bootstrapping and starts the
|
||||
job tracker.
|
||||
|
||||
After the master machine gets up, we can freely add more slave machines to the cluster.
|
||||
The following command add m3.xlarge instances to the cluster.
|
||||
```bash
|
||||
./yarn-ec2 -k mykey -i mypem.pem -t m3.xlarge -s 2 addslave xgboost
|
||||
```
|
||||
We can also choose to add two spot instances
|
||||
```bash
|
||||
./yarn-ec2 -k mykey -i mypem.pem -t m3.xlarge -s 2 addspot xgboost
|
||||
```
|
||||
The slave machines will start up, bootstrap and report to the master.
|
||||
You can check if the slave machines are connected by clicking on the Nodes link on the job tracker.
|
||||
Or simply type the following URL (replace DNS ith the master DNS)
|
||||
```
|
||||
ec2-xx-xx-xx.us-west-2.compute.amazonaws.com:8088/cluster/nodes
|
||||
```
|
||||
|
||||
One thing we should note is that not all the links in the job tracker work.
|
||||
This is due to that many of them use the private IP of AWS, which can only be accessed by EC2.
|
||||
We can use ssh proxy to access these packages.
|
||||
Now that we have set up a cluster with one master and two slaves, we are ready to run the experiment.
|
||||
|
||||
|
||||
Build XGBoost with S3
|
||||
---------------------
|
||||
We can log into the master machine by the following command.
|
||||
```bash
|
||||
./yarn-ec2 -k mykey -i mypem.pem login xgboost
|
||||
```
|
||||
|
||||
We will be using S3 to host the data and the result model, so the data won't get lost after the cluster shutdown.
|
||||
To do so, we will need to build xgboost with S3 support. The only thing we need to do is to set ```USE_S3```
|
||||
variable to be true. This can be achieved by the following command.
|
||||
|
||||
```bash
|
||||
git clone --recursive https://github.com/dmlc/xgboost
|
||||
cd xgboost
|
||||
cp make/config.mk config.mk
|
||||
echo "USE_S3=1" >> config.mk
|
||||
make -j4
|
||||
```
|
||||
Now we have built the XGBoost with S3 support. You can also enable HDFS support if you plan to store data on HDFS by turning on ```USE_HDFS``` option.
|
||||
|
||||
XGBoost also relies on the environment variable to access S3, so you will need to add the following two lines to `~/.bashrc` (replacing the strings with the correct ones)
|
||||
on the master machine as well.
|
||||
|
||||
```bash
|
||||
export AWS_ACCESS_KEY_ID=AKIAIOSFODNN7EXAMPLE
|
||||
export AWS_SECRET_ACCESS_KEY=wJalrXUtnFEMI/K7MDENG/bPxRfiCYEXAMPLEKEY
|
||||
export BUCKET=mybucket
|
||||
```
|
||||
|
||||
Host the Data on S3
|
||||
-------------------
|
||||
In this example, we will copy the example dataset in xgboost to the S3 bucket as input.
|
||||
In normal usecases, the dataset is usually created from existing distributed processing pipeline.
|
||||
We can use [s3cmd](http://s3tools.org/s3cmd) to copy the data into mybucket (replace ${BUCKET} with the real bucket name).
|
||||
|
||||
```bash
|
||||
cd xgboost
|
||||
s3cmd put demo/data/agaricus.txt.train s3://${BUCKET}/xgb-demo/train/
|
||||
s3cmd put demo/data/agaricus.txt.test s3://${BUCKET}/xgb-demo/test/
|
||||
```
|
||||
|
||||
Submit the Jobs
|
||||
---------------
|
||||
Now everything is ready, we can submit the xgboost distributed job to the YARN cluster.
|
||||
We will use the [dmlc-submit](https://github.com/dmlc/dmlc-core/tree/master/tracker) script to submit the job.
|
||||
|
||||
Now we can run the following script in the distributed training folder (replace ${BUCKET} with the real bucket name)
|
||||
```bash
|
||||
cd xgboost/demo/distributed-training
|
||||
# Use dmlc-submit to submit the job.
|
||||
../../dmlc-core/tracker/dmlc-submit --cluster=yarn --num-workers=2 --worker-cores=2\
|
||||
../../xgboost mushroom.aws.conf nthread=2\
|
||||
data=s3://${BUCKET}/xgb-demo/train\
|
||||
eval[test]=s3://${BUCKET}/xgb-demo/test\
|
||||
model_dir=s3://${BUCKET}/xgb-demo/model
|
||||
```
|
||||
All the configurations such as ```data``` and ```model_dir``` can also be directly written into the configuration file.
|
||||
Note that we only specified the folder path to the file, instead of the file name.
|
||||
XGBoost will read in all the files in that folder as the training and evaluation data.
|
||||
|
||||
In this command, we are using two workers, and each worker uses two running threads.
|
||||
XGBoost can benefit from using multiple cores in each worker.
|
||||
A common choice of working cores can range from 4 to 8.
|
||||
The trained model will be saved into the specified model folder. You can browse the model folder.
|
||||
```
|
||||
s3cmd ls s3://${BUCKET}/xgb-demo/model/
|
||||
```
|
||||
|
||||
The following is an example output from distributed training.
|
||||
```
|
||||
16/02/26 05:41:59 INFO dmlc.Client: jobname=DMLC[nworker=2]:xgboost,username=ubuntu
|
||||
16/02/26 05:41:59 INFO dmlc.Client: Submitting application application_1456461717456_0015
|
||||
16/02/26 05:41:59 INFO impl.YarnClientImpl: Submitted application application_1456461717456_0015
|
||||
2016-02-26 05:42:05,230 INFO @tracker All of 2 nodes getting started
|
||||
2016-02-26 05:42:14,027 INFO [05:42:14] [0] test-error:0.016139 train-error:0.014433
|
||||
2016-02-26 05:42:14,186 INFO [05:42:14] [1] test-error:0.000000 train-error:0.001228
|
||||
2016-02-26 05:42:14,947 INFO @tracker All nodes finishes job
|
||||
2016-02-26 05:42:14,948 INFO @tracker 9.71754479408 secs between node start and job finish
|
||||
Application application_1456461717456_0015 finished with state FINISHED at 1456465335961
|
||||
```
|
||||
|
||||
Analyze the Model
|
||||
-----------------
|
||||
After the model is trained, we can analyse the learnt model and use it for future prediction tasks.
|
||||
XGBoost is a portable framework, meaning the models in all platforms are ***exchangeable***.
|
||||
This means we can load the trained model in python/R/Julia and take benefit of data science pipelines
|
||||
in these languages to do model analysis and prediction.
|
||||
|
||||
For example, you can use [this IPython notebook](https://github.com/dmlc/xgboost/tree/master/demo/distributed-training/plot_model.ipynb)
|
||||
to plot feature importance and visualize the learnt model.
|
||||
|
||||
Troubleshooting
|
||||
----------------
|
||||
|
||||
If you encounter a problem, the best way might be to use the following command
|
||||
to get logs of stdout and stderr of the containers and check what causes the problem.
|
||||
```
|
||||
yarn logs -applicationId yourAppId
|
||||
```
|
||||
|
||||
Future Directions
|
||||
-----------------
|
||||
You have learned to use distributed XGBoost on YARN in this tutorial.
|
||||
XGBoost is a portable and scalable framework for gradient boosting.
|
||||
You can check out more examples and resources in the [resources page](https://github.com/dmlc/xgboost/blob/master/demo/README.md).
|
||||
|
||||
The project goal is to make the best scalable machine learning solution available to all platforms.
|
||||
The API is designed to be able to portable, and the same code can also run on other platforms such as MPI and SGE.
|
||||
XGBoost is actively evolving and we are working on even more exciting features
|
||||
such as distributed xgboost python/R package. Checkout [RoadMap](https://github.com/dmlc/xgboost/issues/873) for
|
||||
more details and you are more than welcomed to contribute to the project.
|
||||
216
doc/tutorials/aws_yarn.rst
Normal file
216
doc/tutorials/aws_yarn.rst
Normal file
@@ -0,0 +1,216 @@
|
||||
###############################
|
||||
Distributed XGBoost YARN on AWS
|
||||
###############################
|
||||
This is a step-by-step tutorial on how to setup and run distributed `XGBoost <https://github.com/dmlc/xgboost>`_
|
||||
on an AWS EC2 cluster. Distributed XGBoost runs on various platforms such as MPI, SGE and Hadoop YARN.
|
||||
In this tutorial, we use YARN as an example since this is a widely used solution for distributed computing.
|
||||
|
||||
.. note:: XGBoost on Spark
|
||||
|
||||
If you are preprocessing training data with Spark, you may want to look at `XGBoost4J-Spark <https://xgboost.ai/2016/10/26/a-full-integration-of-xgboost-and-spark.html>`_, which supports distributed training on Resilient Distributed Dataset (RDD).
|
||||
|
||||
************
|
||||
Prerequisite
|
||||
************
|
||||
We need to get a `AWS key-pair <http://docs.aws.amazon.com/AWSEC2/latest/UserGuide/ec2-key-pairs.html>`_
|
||||
to access the AWS services. Let us assume that we are using a key ``mykey`` and the corresponding permission file ``mypem.pem``.
|
||||
|
||||
We also need `AWS credentials <https://docs.aws.amazon.com/cli/latest/userguide/cli-chap-getting-started.html>`_,
|
||||
which includes an ``ACCESS_KEY_ID`` and a ``SECRET_ACCESS_KEY``.
|
||||
|
||||
Finally, we will need a S3 bucket to host the data and the model, ``s3://mybucket/``
|
||||
|
||||
***************************
|
||||
Setup a Hadoop YARN Cluster
|
||||
***************************
|
||||
This sections shows how to start a Hadoop YARN cluster from scratch.
|
||||
You can skip this step if you have already have one.
|
||||
We will be using `yarn-ec2 <https://github.com/tqchen/yarn-ec2>`_ to start the cluster.
|
||||
|
||||
We can first clone the yarn-ec2 script by the following command.
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
git clone https://github.com/tqchen/yarn-ec2
|
||||
|
||||
To use the script, we must set the environment variables ``AWS_ACCESS_KEY_ID`` and
|
||||
``AWS_SECRET_ACCESS_KEY`` properly. This can be done by adding the following two lines in
|
||||
``~/.bashrc`` (replacing the strings with the correct ones)
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
export AWS_ACCESS_KEY_ID=[your access ID]
|
||||
export AWS_SECRET_ACCESS_KEY=[your secret access key]
|
||||
|
||||
Now we can launch a master machine of the cluster from EC2:
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
./yarn-ec2 -k mykey -i mypem.pem launch xgboost
|
||||
|
||||
Wait a few mininutes till the master machine gets up.
|
||||
|
||||
After the master machine gets up, we can query the public DNS of the master machine using the following command.
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
./yarn-ec2 -k mykey -i mypem.pem get-master xgboost
|
||||
|
||||
It will show the public DNS of the master machine like ``ec2-xx-xx-xx.us-west-2.compute.amazonaws.com``
|
||||
Now we can open the browser, and type (replace the DNS with the master DNS)
|
||||
|
||||
.. code-block:: none
|
||||
|
||||
ec2-xx-xx-xx.us-west-2.compute.amazonaws.com:8088
|
||||
|
||||
This will show the job tracker of the YARN cluster. Note that we may have to wait a few minutes before the master finishes bootstrapping and starts the
|
||||
job tracker.
|
||||
|
||||
After the master machine gets up, we can freely add more slave machines to the cluster.
|
||||
The following command add m3.xlarge instances to the cluster.
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
./yarn-ec2 -k mykey -i mypem.pem -t m3.xlarge -s 2 addslave xgboost
|
||||
|
||||
We can also choose to add two spot instances
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
./yarn-ec2 -k mykey -i mypem.pem -t m3.xlarge -s 2 addspot xgboost
|
||||
|
||||
The slave machines will start up, bootstrap and report to the master.
|
||||
You can check if the slave machines are connected by clicking on the Nodes link on the job tracker.
|
||||
Or simply type the following URL (replace DNS ith the master DNS)
|
||||
|
||||
.. code-block:: none
|
||||
|
||||
ec2-xx-xx-xx.us-west-2.compute.amazonaws.com:8088/cluster/nodes
|
||||
|
||||
One thing we should note is that not all the links in the job tracker work.
|
||||
This is due to that many of them use the private IP of AWS, which can only be accessed by EC2.
|
||||
We can use ssh proxy to access these packages.
|
||||
Now that we have set up a cluster with one master and two slaves, we are ready to run the experiment.
|
||||
|
||||
*********************
|
||||
Build XGBoost with S3
|
||||
*********************
|
||||
We can log into the master machine by the following command.
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
./yarn-ec2 -k mykey -i mypem.pem login xgboost
|
||||
|
||||
We will be using S3 to host the data and the result model, so the data won't get lost after the cluster shutdown.
|
||||
To do so, we will need to build XGBoost with S3 support. The only thing we need to do is to set ``USE_S3``
|
||||
variable to be true. This can be achieved by the following command.
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
git clone --recursive https://github.com/dmlc/xgboost
|
||||
cd xgboost
|
||||
cp make/config.mk config.mk
|
||||
echo "USE_S3=1" >> config.mk
|
||||
make -j4
|
||||
|
||||
Now we have built the XGBoost with S3 support. You can also enable HDFS support if you plan to store data on HDFS by turning on ``USE_HDFS`` option.
|
||||
XGBoost also relies on the environment variable to access S3, so you will need to add the following two lines to ``~/.bashrc`` (replacing the strings with the correct ones)
|
||||
on the master machine as well.
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
export AWS_ACCESS_KEY_ID=AKIAIOSFODNN7EXAMPLE
|
||||
export AWS_SECRET_ACCESS_KEY=wJalrXUtnFEMI/K7MDENG/bPxRfiCYEXAMPLEKEY
|
||||
export BUCKET=mybucket
|
||||
|
||||
*******************
|
||||
Host the Data on S3
|
||||
*******************
|
||||
In this example, we will copy the example dataset in XGBoost to the S3 bucket as input.
|
||||
In normal usecases, the dataset is usually created from existing distributed processing pipeline.
|
||||
We can use `s3cmd <http://s3tools.org/s3cmd>`_ to copy the data into mybucket (replace ``${BUCKET}`` with the real bucket name).
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
cd xgboost
|
||||
s3cmd put demo/data/agaricus.txt.train s3://${BUCKET}/xgb-demo/train/
|
||||
s3cmd put demo/data/agaricus.txt.test s3://${BUCKET}/xgb-demo/test/
|
||||
|
||||
***************
|
||||
Submit the Jobs
|
||||
***************
|
||||
Now everything is ready, we can submit the XGBoost distributed job to the YARN cluster.
|
||||
We will use the `dmlc-submit <https://github.com/dmlc/dmlc-core/tree/master/tracker>`_ script to submit the job.
|
||||
|
||||
Now we can run the following script in the distributed training folder (replace ``${BUCKET}`` with the real bucket name)
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
cd xgboost/demo/distributed-training
|
||||
# Use dmlc-submit to submit the job.
|
||||
../../dmlc-core/tracker/dmlc-submit --cluster=yarn --num-workers=2 --worker-cores=2\
|
||||
../../xgboost mushroom.aws.conf nthread=2\
|
||||
data=s3://${BUCKET}/xgb-demo/train\
|
||||
eval[test]=s3://${BUCKET}/xgb-demo/test\
|
||||
model_dir=s3://${BUCKET}/xgb-demo/model
|
||||
|
||||
All the configurations such as ``data`` and ``model_dir`` can also be directly written into the configuration file.
|
||||
Note that we only specified the folder path to the file, instead of the file name.
|
||||
XGBoost will read in all the files in that folder as the training and evaluation data.
|
||||
|
||||
In this command, we are using two workers, and each worker uses two running threads.
|
||||
XGBoost can benefit from using multiple cores in each worker.
|
||||
A common choice of working cores can range from 4 to 8.
|
||||
The trained model will be saved into the specified model folder. You can browse the model folder.
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
s3cmd ls s3://${BUCKET}/xgb-demo/model/
|
||||
|
||||
The following is an example output from distributed training.
|
||||
|
||||
.. code-block:: none
|
||||
|
||||
16/02/26 05:41:59 INFO dmlc.Client: jobname=DMLC[nworker=2]:xgboost,username=ubuntu
|
||||
16/02/26 05:41:59 INFO dmlc.Client: Submitting application application_1456461717456_0015
|
||||
16/02/26 05:41:59 INFO impl.YarnClientImpl: Submitted application application_1456461717456_0015
|
||||
2016-02-26 05:42:05,230 INFO @tracker All of 2 nodes getting started
|
||||
2016-02-26 05:42:14,027 INFO [05:42:14] [0] test-error:0.016139 train-error:0.014433
|
||||
2016-02-26 05:42:14,186 INFO [05:42:14] [1] test-error:0.000000 train-error:0.001228
|
||||
2016-02-26 05:42:14,947 INFO @tracker All nodes finishes job
|
||||
2016-02-26 05:42:14,948 INFO @tracker 9.71754479408 secs between node start and job finish
|
||||
Application application_1456461717456_0015 finished with state FINISHED at 1456465335961
|
||||
|
||||
*****************
|
||||
Analyze the Model
|
||||
*****************
|
||||
After the model is trained, we can analyse the learnt model and use it for future prediction tasks.
|
||||
XGBoost is a portable framework, meaning the models in all platforms are *exchangeable*.
|
||||
This means we can load the trained model in python/R/Julia and take benefit of data science pipelines
|
||||
in these languages to do model analysis and prediction.
|
||||
|
||||
For example, you can use `this IPython notebook <https://github.com/dmlc/xgboost/tree/master/demo/distributed-training/plot_model.ipynb>`_
|
||||
to plot feature importance and visualize the learnt model.
|
||||
|
||||
***************
|
||||
Troubleshooting
|
||||
***************
|
||||
|
||||
If you encounter a problem, the best way might be to use the following command
|
||||
to get logs of stdout and stderr of the containers and check what causes the problem.
|
||||
|
||||
.. code-block:: bash
|
||||
|
||||
yarn logs -applicationId yourAppId
|
||||
|
||||
*****************
|
||||
Future Directions
|
||||
*****************
|
||||
You have learned to use distributed XGBoost on YARN in this tutorial.
|
||||
XGBoost is a portable and scalable framework for gradient boosting.
|
||||
You can check out more examples and resources in the `resources page <https://github.com/dmlc/xgboost/blob/master/demo/README.md>`_.
|
||||
|
||||
The project goal is to make the best scalable machine learning solution available to all platforms.
|
||||
The API is designed to be able to portable, and the same code can also run on other platforms such as MPI and SGE.
|
||||
XGBoost is actively evolving and we are working on even more exciting features
|
||||
such as distributed XGBoost python/R package.
|
||||
@@ -1,101 +0,0 @@
|
||||
DART booster
|
||||
============
|
||||
[XGBoost](https://github.com/dmlc/xgboost) mostly combines a huge number of regression trees with a small learning rate.
|
||||
In this situation, trees added early are significant and trees added late are unimportant.
|
||||
|
||||
Vinayak and Gilad-Bachrach proposed a new method to add dropout techniques from the deep neural net community to boosted trees, and reported better results in some situations.
|
||||
|
||||
This is a instruction of new tree booster `dart`.
|
||||
|
||||
Original paper
|
||||
--------------
|
||||
Rashmi Korlakai Vinayak, Ran Gilad-Bachrach. "DART: Dropouts meet Multiple Additive Regression Trees." [JMLR](http://www.jmlr.org/proceedings/papers/v38/korlakaivinayak15.pdf)
|
||||
|
||||
Features
|
||||
--------
|
||||
- Drop trees in order to solve the over-fitting.
|
||||
- Trivial trees (to correct trivial errors) may be prevented.
|
||||
|
||||
Because of the randomness introduced in the training, expect the following few differences:
|
||||
- Training can be slower than `gbtree` because the random dropout prevents usage of the prediction buffer.
|
||||
- The early stop might not be stable, due to the randomness.
|
||||
|
||||
How it works
|
||||
------------
|
||||
- In ``$ m $``th training round, suppose ``$ k $`` trees are selected to be dropped.
|
||||
- Let ``$ D = \sum_{i \in \mathbf{K}} F_i $`` be the leaf scores of dropped trees and ``$ F_m = \eta \tilde{F}_m $`` be the leaf scores of a new tree.
|
||||
- The objective function is as follows:
|
||||
```math
|
||||
\mathrm{Obj}
|
||||
= \sum_{j=1}^n L \left( y_j, \hat{y}_j^{m-1} - D_j + \tilde{F}_m \right)
|
||||
+ \Omega \left( \tilde{F}_m \right).
|
||||
```
|
||||
- ``$ D $`` and ``$ F_m $`` are overshooting, so using scale factor
|
||||
```math
|
||||
\hat{y}_j^m = \sum_{i \not\in \mathbf{K}} F_i + a \left( \sum_{i \in \mathbf{K}} F_i + b F_m \right) .
|
||||
```
|
||||
|
||||
Parameters
|
||||
----------
|
||||
### booster
|
||||
* `dart`
|
||||
|
||||
This booster inherits `gbtree`, so `dart` has also `eta`, `gamma`, `max_depth` and so on.
|
||||
|
||||
Additional parameters are noted below.
|
||||
|
||||
### sample_type
|
||||
type of sampling algorithm.
|
||||
* `uniform`: (default) dropped trees are selected uniformly.
|
||||
* `weighted`: dropped trees are selected in proportion to weight.
|
||||
|
||||
### normalize_type
|
||||
type of normalization algorithm.
|
||||
* `tree`: (default) New trees have the same weight of each of dropped trees.
|
||||
```math
|
||||
a \left( \sum_{i \in \mathbf{K}} F_i + \frac{1}{k} F_m \right)
|
||||
&= a \left( \sum_{i \in \mathbf{K}} F_i + \frac{\eta}{k} \tilde{F}_m \right) \\
|
||||
&\sim a \left( 1 + \frac{\eta}{k} \right) D \\
|
||||
&= a \frac{k + \eta}{k} D = D , \\
|
||||
&\quad a = \frac{k}{k + \eta} .
|
||||
```
|
||||
|
||||
* `forest`: New trees have the same weight of sum of dropped trees (forest).
|
||||
```math
|
||||
a \left( \sum_{i \in \mathbf{K}} F_i + F_m \right)
|
||||
&= a \left( \sum_{i \in \mathbf{K}} F_i + \eta \tilde{F}_m \right) \\
|
||||
&\sim a \left( 1 + \eta \right) D \\
|
||||
&= a (1 + \eta) D = D , \\
|
||||
&\quad a = \frac{1}{1 + \eta} .
|
||||
```
|
||||
|
||||
### rate_drop
|
||||
dropout rate.
|
||||
- range: [0.0, 1.0]
|
||||
|
||||
### skip_drop
|
||||
probability of skipping dropout.
|
||||
- If a dropout is skipped, new trees are added in the same manner as gbtree.
|
||||
- range: [0.0, 1.0]
|
||||
|
||||
Sample Script
|
||||
-------------
|
||||
```python
|
||||
import xgboost as xgb
|
||||
# read in data
|
||||
dtrain = xgb.DMatrix('demo/data/agaricus.txt.train')
|
||||
dtest = xgb.DMatrix('demo/data/agaricus.txt.test')
|
||||
# specify parameters via map
|
||||
param = {'booster': 'dart',
|
||||
'max_depth': 5, 'learning_rate': 0.1,
|
||||
'objective': 'binary:logistic', 'silent': True,
|
||||
'sample_type': 'uniform',
|
||||
'normalize_type': 'tree',
|
||||
'rate_drop': 0.1,
|
||||
'skip_drop': 0.5}
|
||||
num_round = 50
|
||||
bst = xgb.train(param, dtrain, num_round)
|
||||
# make prediction
|
||||
# ntree_limit must not be 0
|
||||
preds = bst.predict(dtest, ntree_limit=num_round)
|
||||
```
|
||||
113
doc/tutorials/dart.rst
Normal file
113
doc/tutorials/dart.rst
Normal file
@@ -0,0 +1,113 @@
|
||||
############
|
||||
DART booster
|
||||
############
|
||||
XGBoost mostly combines a huge number of regression trees with a small learning rate.
|
||||
In this situation, trees added early are significant and trees added late are unimportant.
|
||||
|
||||
Vinayak and Gilad-Bachrach proposed a new method to add dropout techniques from the deep neural net community to boosted trees, and reported better results in some situations.
|
||||
|
||||
This is a instruction of new tree booster ``dart``.
|
||||
|
||||
**************
|
||||
Original paper
|
||||
**************
|
||||
Rashmi Korlakai Vinayak, Ran Gilad-Bachrach. "DART: Dropouts meet Multiple Additive Regression Trees." `JMLR <http://www.jmlr.org/proceedings/papers/v38/korlakaivinayak15.pdf>`_.
|
||||
|
||||
********
|
||||
Features
|
||||
********
|
||||
- Drop trees in order to solve the over-fitting.
|
||||
|
||||
- Trivial trees (to correct trivial errors) may be prevented.
|
||||
|
||||
Because of the randomness introduced in the training, expect the following few differences:
|
||||
|
||||
- Training can be slower than ``gbtree`` because the random dropout prevents usage of the prediction buffer.
|
||||
- The early stop might not be stable, due to the randomness.
|
||||
|
||||
************
|
||||
How it works
|
||||
************
|
||||
- In :math:`m`-th training round, suppose :math:`k` trees are selected to be dropped.
|
||||
- Let :math:`D = \sum_{i \in \mathbf{K}} F_i` be the leaf scores of dropped trees and :math:`F_m = \eta \tilde{F}_m` be the leaf scores of a new tree.
|
||||
- The objective function is as follows:
|
||||
|
||||
.. math::
|
||||
|
||||
\mathrm{Obj}
|
||||
= \sum_{j=1}^n L \left( y_j, \hat{y}_j^{m-1} - D_j + \tilde{F}_m \right)
|
||||
+ \Omega \left( \tilde{F}_m \right).
|
||||
|
||||
- :math:`D` and :math:`F_m` are overshooting, so using scale factor
|
||||
|
||||
.. math::
|
||||
|
||||
\hat{y}_j^m = \sum_{i \not\in \mathbf{K}} F_i + a \left( \sum_{i \in \mathbf{K}} F_i + b F_m \right) .
|
||||
|
||||
**********
|
||||
Parameters
|
||||
**********
|
||||
|
||||
The booster ``dart`` inherits ``gbtree`` booster, so it supports all parameters that ``gbtree`` does, such as ``eta``, ``gamma``, ``max_depth`` etc.
|
||||
|
||||
Additional parameters are noted below:
|
||||
|
||||
* ``sample_type``: type of sampling algorithm.
|
||||
|
||||
- ``uniform``: (default) dropped trees are selected uniformly.
|
||||
- ``weighted``: dropped trees are selected in proportion to weight.
|
||||
|
||||
* ``normalize_type``: type of normalization algorithm.
|
||||
|
||||
- ``tree``: (default) New trees have the same weight of each of dropped trees.
|
||||
|
||||
.. math::
|
||||
|
||||
a \left( \sum_{i \in \mathbf{K}} F_i + \frac{1}{k} F_m \right)
|
||||
&= a \left( \sum_{i \in \mathbf{K}} F_i + \frac{\eta}{k} \tilde{F}_m \right) \\
|
||||
&\sim a \left( 1 + \frac{\eta}{k} \right) D \\
|
||||
&= a \frac{k + \eta}{k} D = D , \\
|
||||
&\quad a = \frac{k}{k + \eta}
|
||||
|
||||
- ``forest``: New trees have the same weight of sum of dropped trees (forest).
|
||||
|
||||
.. math::
|
||||
|
||||
a \left( \sum_{i \in \mathbf{K}} F_i + F_m \right)
|
||||
&= a \left( \sum_{i \in \mathbf{K}} F_i + \eta \tilde{F}_m \right) \\
|
||||
&\sim a \left( 1 + \eta \right) D \\
|
||||
&= a (1 + \eta) D = D , \\
|
||||
&\quad a = \frac{1}{1 + \eta} .
|
||||
|
||||
* ``rate_drop``: dropout rate.
|
||||
|
||||
- range: [0.0, 1.0]
|
||||
|
||||
* ``skip_drop``: probability of skipping dropout.
|
||||
|
||||
- If a dropout is skipped, new trees are added in the same manner as gbtree.
|
||||
- range: [0.0, 1.0]
|
||||
|
||||
*************
|
||||
Sample Script
|
||||
*************
|
||||
|
||||
.. code-block:: python
|
||||
|
||||
import xgboost as xgb
|
||||
# read in data
|
||||
dtrain = xgb.DMatrix('demo/data/agaricus.txt.train')
|
||||
dtest = xgb.DMatrix('demo/data/agaricus.txt.test')
|
||||
# specify parameters via map
|
||||
param = {'booster': 'dart',
|
||||
'max_depth': 5, 'learning_rate': 0.1,
|
||||
'objective': 'binary:logistic', 'silent': True,
|
||||
'sample_type': 'uniform',
|
||||
'normalize_type': 'tree',
|
||||
'rate_drop': 0.1,
|
||||
'skip_drop': 0.5}
|
||||
num_round = 50
|
||||
bst = xgb.train(param, dtrain, num_round)
|
||||
# make prediction
|
||||
# ntree_limit must not be 0
|
||||
preds = bst.predict(dtest, ntree_limit=num_round)
|
||||
51
doc/tutorials/external_memory.rst
Normal file
51
doc/tutorials/external_memory.rst
Normal file
@@ -0,0 +1,51 @@
|
||||
############################################
|
||||
Using XGBoost External Memory Version (beta)
|
||||
############################################
|
||||
There is no big difference between using external memory version and in-memory version.
|
||||
The only difference is the filename format.
|
||||
|
||||
The external memory version takes in the following filename format:
|
||||
|
||||
.. code-block:: none
|
||||
|
||||
filename#cacheprefix
|
||||
|
||||
The ``filename`` is the normal path to libsvm file you want to load in, and ``cacheprefix`` is a
|
||||
path to a cache file that XGBoost will use for external memory cache.
|
||||
|
||||
The following code was extracted from `demo/guide-python/external_memory.py <https://github.com/dmlc/xgboost/blob/master/demo/guide-python/external_memory.py>`_:
|
||||
|
||||
.. code-block:: python
|
||||
|
||||
dtrain = xgb.DMatrix('../data/agaricus.txt.train#dtrain.cache')
|
||||
|
||||
You can find that there is additional ``#dtrain.cache`` following the libsvm file, this is the name of cache file.
|
||||
For CLI version, simply add the cache suffix, e.g. ``"../data/agaricus.txt.train#dtrain.cache"``.
|
||||
|
||||
****************
|
||||
Performance Note
|
||||
****************
|
||||
* the parameter ``nthread`` should be set to number of **physical** cores
|
||||
|
||||
- Most modern CPUs use hyperthreading, which means a 4 core CPU may carry 8 threads
|
||||
- Set ``nthread`` to be 4 for maximum performance in such case
|
||||
|
||||
*******************
|
||||
Distributed Version
|
||||
*******************
|
||||
The external memory mode naturally works on distributed version, you can simply set path like
|
||||
|
||||
.. code-block:: none
|
||||
|
||||
data = "hdfs://path-to-data/#dtrain.cache"
|
||||
|
||||
XGBoost will cache the data to the local position. When you run on YARN, the current folder is temporal
|
||||
so that you can directly use ``dtrain.cache`` to cache to current folder.
|
||||
|
||||
**********
|
||||
Usage Note
|
||||
**********
|
||||
* This is a experimental version
|
||||
* Currently only importing from libsvm format is supported
|
||||
|
||||
- Contribution of ingestion from other common external memory data source is welcomed
|
||||
@@ -1,10 +0,0 @@
|
||||
# XGBoost Tutorials
|
||||
|
||||
This section contains official tutorials inside XGBoost package.
|
||||
See [Awesome XGBoost](https://github.com/dmlc/xgboost/tree/master/demo) for links to more resources.
|
||||
|
||||
## Contents
|
||||
- [Introduction to Boosted Trees](../model.md)
|
||||
- [Distributed XGBoost YARN on AWS](aws_yarn.md)
|
||||
- [DART booster](dart.md)
|
||||
- [Monotonic Constraints](monotonic.md)
|
||||
19
doc/tutorials/index.rst
Normal file
19
doc/tutorials/index.rst
Normal file
@@ -0,0 +1,19 @@
|
||||
#################
|
||||
XGBoost Tutorials
|
||||
#################
|
||||
|
||||
This section contains official tutorials inside XGBoost package.
|
||||
See `Awesome XGBoost <https://github.com/dmlc/xgboost/tree/master/demo>`_ for more resources.
|
||||
|
||||
.. toctree::
|
||||
:maxdepth: 1
|
||||
:caption: Contents:
|
||||
|
||||
model
|
||||
aws_yarn
|
||||
dart
|
||||
monotonic
|
||||
input_format
|
||||
param_tuning
|
||||
external_memory
|
||||
|
||||
112
doc/tutorials/input_format.rst
Normal file
112
doc/tutorials/input_format.rst
Normal file
@@ -0,0 +1,112 @@
|
||||
############################
|
||||
Text Input Format of DMatrix
|
||||
############################
|
||||
|
||||
******************
|
||||
Basic Input Format
|
||||
******************
|
||||
XGBoost currently supports two text formats for ingesting data: LibSVM and CSV. The rest of this document will describe the LibSVM format. (See `this Wikipedia article <https://en.wikipedia.org/wiki/Comma-separated_values>`_ for a description of the CSV format.)
|
||||
|
||||
For training or predicting, XGBoost takes an instance file with the format as below:
|
||||
|
||||
.. code-block:: none
|
||||
:caption: ``train.txt``
|
||||
|
||||
1 101:1.2 102:0.03
|
||||
0 1:2.1 10001:300 10002:400
|
||||
0 0:1.3 1:0.3
|
||||
1 0:0.01 1:0.3
|
||||
0 0:0.2 1:0.3
|
||||
|
||||
Each line represent a single instance, and in the first line '1' is the instance label, '101' and '102' are feature indices, '1.2' and '0.03' are feature values. In the binary classification case, '1' is used to indicate positive samples, and '0' is used to indicate negative samples. We also support probability values in [0,1] as label, to indicate the probability of the instance being positive.
|
||||
|
||||
******************************************
|
||||
Auxiliary Files for Additional Information
|
||||
******************************************
|
||||
**Note: all information below is applicable only to single-node version of the package.** If you'd like to perform distributed training with multiple nodes, skip to the section `Embedding additional information inside LibSVM file`_.
|
||||
|
||||
Group Input Format
|
||||
==================
|
||||
For `ranking task <https://github.com/dmlc/xgboost/tree/master/demo/rank>`_, XGBoost supports the group input format. In ranking task, instances are categorized into *query groups* in real world scenarios. For example, in the learning to rank web pages scenario, the web page instances are grouped by their queries. XGBoost requires an file that indicates the group information. For example, if the instance file is the ``train.txt`` shown above, the group file should be named ``train.txt.group`` and be of the following format:
|
||||
|
||||
.. code-block:: none
|
||||
:caption: ``train.txt.group``
|
||||
|
||||
2
|
||||
3
|
||||
|
||||
This means that, the data set contains 5 instances, and the first two instances are in a group and the other three are in another group. The numbers in the group file are actually indicating the number of instances in each group in the instance file in order.
|
||||
At the time of configuration, you do not have to indicate the path of the group file. If the instance file name is ``xxx``, XGBoost will check whether there is a file named ``xxx.group`` in the same directory.
|
||||
|
||||
Instance Weight File
|
||||
====================
|
||||
Instances in the training data may be assigned weights to differentiate relative importance among them. For example, if we provide an instance weight file for the ``train.txt`` file in the example as below:
|
||||
|
||||
.. code-block:: none
|
||||
:caption: ``train.txt.weight``
|
||||
|
||||
1
|
||||
0.5
|
||||
0.5
|
||||
1
|
||||
0.5
|
||||
|
||||
It means that XGBoost will emphasize more on the first and fourth instance (i.e. the positive instances) while training.
|
||||
The configuration is similar to configuring the group information. If the instance file name is ``xxx``, XGBoost will look for a file named ``xxx.weight`` in the same directory. If the file exists, the instance weights will be extracted and used at the time of training.
|
||||
|
||||
.. note:: Binary buffer format and instance weights
|
||||
|
||||
If you choose to save the training data as a binary buffer (using :py:meth:`save_binary() <xgboost.DMatrix.save_binary>`), keep in mind that the resulting binary buffer file will include the instance weights. To update the weights, use the :py:meth:`set_weight() <xgboost.DMatrix.set_weight>` function.
|
||||
|
||||
Initial Margin File
|
||||
===================
|
||||
XGBoost supports providing each instance an initial margin prediction. For example, if we have a initial prediction using logistic regression for ``train.txt`` file, we can create the following file:
|
||||
|
||||
.. code-block:: none
|
||||
:caption: ``train.txt.base_margin``
|
||||
|
||||
-0.4
|
||||
1.0
|
||||
3.4
|
||||
|
||||
XGBoost will take these values as initial margin prediction and boost from that. An important note about base_margin is that it should be margin prediction before transformation, so if you are doing logistic loss, you will need to put in value before logistic transformation. If you are using XGBoost predictor, use ``pred_margin=1`` to output margin values.
|
||||
|
||||
***************************************************
|
||||
Embedding additional information inside LibSVM file
|
||||
***************************************************
|
||||
**This section is applicable to both single- and multiple-node settings.**
|
||||
|
||||
Query ID Columns
|
||||
================
|
||||
This is most useful for `ranking task <https://github.com/dmlc/xgboost/tree/master/demo/rank>`_, where the instances are grouped into query groups. You may embed query group ID for each instance in the LibSVM file by adding a token of form ``qid:xx`` in each row:
|
||||
|
||||
.. code-block:: none
|
||||
:caption: ``train.txt``
|
||||
|
||||
1 qid:1 101:1.2 102:0.03
|
||||
0 qid:1 1:2.1 10001:300 10002:400
|
||||
0 qid:2 0:1.3 1:0.3
|
||||
1 qid:2 0:0.01 1:0.3
|
||||
0 qid:3 0:0.2 1:0.3
|
||||
1 qid:3 3:-0.1 10:-0.3
|
||||
0 qid:3 6:0.2 10:0.15
|
||||
|
||||
Keep in mind the following restrictions:
|
||||
|
||||
* You are not allowed to specify query ID's for some instances but not for others. Either every row is assigned query ID's or none at all.
|
||||
* The rows have to be sorted in ascending order by the query IDs. So, for instance, you may not have one row having large query ID than any of the following rows.
|
||||
|
||||
Instance weights
|
||||
================
|
||||
You may specify instance weights in the LibSVM file by appending each instance label with the corresponding weight in the form of ``[label]:[weight]``, as shown by the following example:
|
||||
|
||||
.. code-block:: none
|
||||
:caption: ``train.txt``
|
||||
|
||||
1:1.0 101:1.2 102:0.03
|
||||
0:0.5 1:2.1 10001:300 10002:400
|
||||
0:0.5 0:1.3 1:0.3
|
||||
1:1.0 0:0.01 1:0.3
|
||||
0:0.5 0:0.2 1:0.3
|
||||
|
||||
where the negative instances are assigned half weights compared to the positive instances.
|
||||
265
doc/tutorials/model.rst
Normal file
265
doc/tutorials/model.rst
Normal file
@@ -0,0 +1,265 @@
|
||||
#############################
|
||||
Introduction to Boosted Trees
|
||||
#############################
|
||||
XGBoost stands for "Extreme Gradient Boosting", where the term "Gradient Boosting" originates from the paper *Greedy Function Approximation: A Gradient Boosting Machine*, by Friedman.
|
||||
This is a tutorial on gradient boosted trees, and most of the content is based on `these slides <http://homes.cs.washington.edu/~tqchen/pdf/BoostedTree.pdf>`_ by Tianqi Chen, the original author of XGBoost.
|
||||
|
||||
The **gradient boosted trees** has been around for a while, and there are a lot of materials on the topic.
|
||||
This tutorial will explain boosted trees in a self-contained and principled way using the elements of supervised learning.
|
||||
We think this explanation is cleaner, more formal, and motivates the model formulation used in XGBoost.
|
||||
|
||||
*******************************
|
||||
Elements of Supervised Learning
|
||||
*******************************
|
||||
XGBoost is used for supervised learning problems, where we use the training data (with multiple features) :math:`x_i` to predict a target variable :math:`y_i`.
|
||||
Before we learn about trees specifically, let us start by reviewing the basic elements in supervised learning.
|
||||
|
||||
Model and Parameters
|
||||
====================
|
||||
The **model** in supervised learning usually refers to the mathematical structure of by which the prediction :math:`y_i` is made from the input :math:`x_i`.
|
||||
A common example is a *linear model*, where the prediction is given as :math:`\hat{y}_i = \sum_j \theta_j x_{ij}`, a linear combination of weighted input features.
|
||||
The prediction value can have different interpretations, depending on the task, i.e., regression or classification.
|
||||
For example, it can be logistic transformed to get the probability of positive class in logistic regression, and it can also be used as a ranking score when we want to rank the outputs.
|
||||
|
||||
The **parameters** are the undetermined part that we need to learn from data. In linear regression problems, the parameters are the coefficients :math:`\theta`.
|
||||
Usually we will use :math:`\theta` to denote the parameters (there are many parameters in a model, our definition here is sloppy).
|
||||
|
||||
Objective Function: Training Loss + Regularization
|
||||
==================================================
|
||||
With judicious choices for :math:`y_i`, we may express a variety of tasks, such as regression, classification, and ranking.
|
||||
The task of **training** the model amounts to finding the best parameters :math:`\theta` that best fit the training data :math:`x_i` and labels :math:`y_i`. In order to train the model, we need to define the **objective function**
|
||||
to measure how well the model fit the training data.
|
||||
|
||||
A salient characteristic of objective functions is that they consist two parts: **training loss** and **regularization term**:
|
||||
|
||||
.. math::
|
||||
|
||||
\text{obj}(\theta) = L(\theta) + \Omega(\theta)
|
||||
|
||||
where :math:`L` is the training loss function, and :math:`\Omega` is the regularization term. The training loss measures how *predictive* our model is with respect to the training data.
|
||||
A common choice of :math:`L` is the *mean squared error*, which is given by
|
||||
|
||||
.. math::
|
||||
|
||||
L(\theta) = \sum_i (y_i-\hat{y}_i)^2
|
||||
|
||||
Another commonly used loss function is logistic loss, to be used for logistic regression:
|
||||
|
||||
.. math::
|
||||
|
||||
L(\theta) = \sum_i[ y_i\ln (1+e^{-\hat{y}_i}) + (1-y_i)\ln (1+e^{\hat{y}_i})]
|
||||
|
||||
The **regularization term** is what people usually forget to add. The regularization term controls the complexity of the model, which helps us to avoid overfitting.
|
||||
This sounds a bit abstract, so let us consider the following problem in the following picture. You are asked to *fit* visually a step function given the input data points
|
||||
on the upper left corner of the image.
|
||||
Which solution among the three do you think is the best fit?
|
||||
|
||||
.. image:: https://raw.githubusercontent.com/dmlc/web-data/master/xgboost/model/step_fit.png
|
||||
:alt: step functions to fit data points, illustrating bias-variance tradeoff
|
||||
|
||||
The correct answer is marked in red. Please consider if this visually seems a reasonable fit to you. The general principle is we want both a *simple* and *predictive* model.
|
||||
The tradeoff between the two is also referred as **bias-variance tradeoff** in machine learning.
|
||||
|
||||
Why introduce the general principle?
|
||||
====================================
|
||||
The elements introduced above form the basic elements of supervised learning, and they are natural building blocks of machine learning toolkits.
|
||||
For example, you should be able to describe the differences and commonalities between gradient boosted trees and random forests.
|
||||
Understanding the process in a formalized way also helps us to understand the objective that we are learning and the reason behind the heuristics such as
|
||||
pruning and smoothing.
|
||||
|
||||
***********************
|
||||
Decision Tree Ensembles
|
||||
***********************
|
||||
Now that we have introduced the elements of supervised learning, let us get started with real trees.
|
||||
To begin with, let us first learn about the model choice of XGBoost: **decision tree ensembles**.
|
||||
The tree ensemble model consists of a set of classification and regression trees (CART). Here's a simple example of a CART
|
||||
that classifies whether someone will like computer games.
|
||||
|
||||
.. image:: https://raw.githubusercontent.com/dmlc/web-data/master/xgboost/model/cart.png
|
||||
:width: 100%
|
||||
:alt: a toy example for CART
|
||||
|
||||
We classify the members of a family into different leaves, and assign them the score on the corresponding leaf.
|
||||
A CART is a bit different from decision trees, in which the leaf only contains decision values. In CART, a real score
|
||||
is associated with each of the leaves, which gives us richer interpretations that go beyond classification.
|
||||
This also allows for a pricipled, unified approach to optimization, as we will see in a later part of this tutorial.
|
||||
|
||||
Usually, a single tree is not strong enough to be used in practice. What is actually used is the ensemble model,
|
||||
which sums the prediction of multiple trees together.
|
||||
|
||||
.. image:: https://raw.githubusercontent.com/dmlc/web-data/master/xgboost/model/twocart.png
|
||||
:width: 100%
|
||||
:alt: a toy example for tree ensemble, consisting of two CARTs
|
||||
|
||||
Here is an example of a tree ensemble of two trees. The prediction scores of each individual tree are summed up to get the final score.
|
||||
If you look at the example, an important fact is that the two trees try to *complement* each other.
|
||||
Mathematically, we can write our model in the form
|
||||
|
||||
.. math::
|
||||
|
||||
\hat{y}_i = \sum_{k=1}^K f_k(x_i), f_k \in \mathcal{F}
|
||||
|
||||
where :math:`K` is the number of trees, :math:`f` is a function in the functional space :math:`\mathcal{F}`, and :math:`\mathcal{F}` is the set of all possible CARTs. The objective function to be optimized is given by
|
||||
|
||||
.. math::
|
||||
|
||||
\text{obj}(\theta) = \sum_i^n l(y_i, \hat{y}_i) + \sum_{k=1}^K \Omega(f_k)
|
||||
|
||||
Now here comes a trick question: what is the *model* used in random forests? Tree ensembles! So random forests and boosted trees are really the same models; the
|
||||
difference arises from how we train them. This means that, if you write a predictive service for tree ensembles, you only need to write one and it should work
|
||||
for both random forests and gradient boosted trees. (See `Treelite <http://treelite.io>`_ for an actual example.) One example of why elements of supervised learning rock.
|
||||
|
||||
*************
|
||||
Tree Boosting
|
||||
*************
|
||||
Now that we introduced the model, let us turn to training: How should we learn the trees?
|
||||
The answer is, as is always for all supervised learning models: *define an objective function and optimize it*!
|
||||
|
||||
Let the following be the objective function (remember it always needs to contain training loss and regularization):
|
||||
|
||||
.. math::
|
||||
|
||||
\text{obj} = \sum_{i=1}^n l(y_i, \hat{y}_i^{(t)}) + \sum_{i=1}^t\Omega(f_i)
|
||||
|
||||
Additive Training
|
||||
=================
|
||||
|
||||
The first question we want to ask: what are the **parameters** of trees?
|
||||
You can find that what we need to learn are those functions :math:`f_i`, each containing the structure
|
||||
of the tree and the leaf scores. Learning tree structure is much harder than traditional optimization problem where you can simply take the gradient.
|
||||
It is intractable to learn all the trees at once.
|
||||
Instead, we use an additive strategy: fix what we have learned, and add one new tree at a time.
|
||||
We write the prediction value at step :math:`t` as :math:`\hat{y}_i^{(t)}`. Then we have
|
||||
|
||||
.. math::
|
||||
|
||||
\hat{y}_i^{(0)} &= 0\\
|
||||
\hat{y}_i^{(1)} &= f_1(x_i) = \hat{y}_i^{(0)} + f_1(x_i)\\
|
||||
\hat{y}_i^{(2)} &= f_1(x_i) + f_2(x_i)= \hat{y}_i^{(1)} + f_2(x_i)\\
|
||||
&\dots\\
|
||||
\hat{y}_i^{(t)} &= \sum_{k=1}^t f_k(x_i)= \hat{y}_i^{(t-1)} + f_t(x_i)
|
||||
|
||||
It remains to ask: which tree do we want at each step? A natural thing is to add the one that optimizes our objective.
|
||||
|
||||
.. math::
|
||||
|
||||
\text{obj}^{(t)} & = \sum_{i=1}^n l(y_i, \hat{y}_i^{(t)}) + \sum_{i=1}^t\Omega(f_i) \\
|
||||
& = \sum_{i=1}^n l(y_i, \hat{y}_i^{(t-1)} + f_t(x_i)) + \Omega(f_t) + \mathrm{constant}
|
||||
|
||||
If we consider using mean squared error (MSE) as our loss function, the objective becomes
|
||||
|
||||
.. math::
|
||||
|
||||
\text{obj}^{(t)} & = \sum_{i=1}^n (y_i - (\hat{y}_i^{(t-1)} + f_t(x_i)))^2 + \sum_{i=1}^t\Omega(f_i) \\
|
||||
& = \sum_{i=1}^n [2(\hat{y}_i^{(t-1)} - y_i)f_t(x_i) + f_t(x_i)^2] + \Omega(f_t) + \mathrm{constant}
|
||||
|
||||
The form of MSE is friendly, with a first order term (usually called the residual) and a quadratic term.
|
||||
For other losses of interest (for example, logistic loss), it is not so easy to get such a nice form.
|
||||
So in the general case, we take the *Taylor expansion of the loss function up to the second order*:
|
||||
|
||||
.. math::
|
||||
|
||||
\text{obj}^{(t)} = \sum_{i=1}^n [l(y_i, \hat{y}_i^{(t-1)}) + g_i f_t(x_i) + \frac{1}{2} h_i f_t^2(x_i)] + \Omega(f_t) + \mathrm{constant}
|
||||
|
||||
where the :math:`g_i` and :math:`h_i` are defined as
|
||||
|
||||
.. math::
|
||||
|
||||
g_i &= \partial_{\hat{y}_i^{(t-1)}} l(y_i, \hat{y}_i^{(t-1)})\\
|
||||
h_i &= \partial_{\hat{y}_i^{(t-1)}}^2 l(y_i, \hat{y}_i^{(t-1)})
|
||||
|
||||
After we remove all the constants, the specific objective at step :math:`t` becomes
|
||||
|
||||
.. math::
|
||||
|
||||
\sum_{i=1}^n [g_i f_t(x_i) + \frac{1}{2} h_i f_t^2(x_i)] + \Omega(f_t)
|
||||
|
||||
This becomes our optimization goal for the new tree. One important advantage of this definition is that
|
||||
the value of the objective function only depends on :math:`g_i` and :math:`h_i`. This is how XGBoost supports custom loss functions.
|
||||
We can optimize every loss function, including logistic regression and pairwise ranking, using exactly
|
||||
the same solver that takes :math:`g_i` and :math:`h_i` as input!
|
||||
|
||||
Model Complexity
|
||||
================
|
||||
We have introduced the training step, but wait, there is one important thing, the **regularization term**!
|
||||
We need to define the complexity of the tree :math:`\Omega(f)`. In order to do so, let us first refine the definition of the tree :math:`f(x)` as
|
||||
|
||||
.. math::
|
||||
|
||||
f_t(x) = w_{q(x)}, w \in R^T, q:R^d\rightarrow \{1,2,\cdots,T\} .
|
||||
|
||||
Here :math:`w` is the vector of scores on leaves, :math:`q` is a function assigning each data point to the corresponding leaf, and :math:`T` is the number of leaves.
|
||||
In XGBoost, we define the complexity as
|
||||
|
||||
.. math::
|
||||
|
||||
\Omega(f) = \gamma T + \frac{1}{2}\lambda \sum_{j=1}^T w_j^2
|
||||
|
||||
Of course, there is more than one way to define the complexity, but this one works well in practice. The regularization is one part most tree packages treat
|
||||
less carefully, or simply ignore. This was because the traditional treatment of tree learning only emphasized improving impurity, while the complexity control was left to heuristics.
|
||||
By defining it formally, we can get a better idea of what we are learning and obtain models that perform well in the wild.
|
||||
|
||||
The Structure Score
|
||||
===================
|
||||
Here is the magical part of the derivation. After re-formulating the tree model, we can write the objective value with the :math:`t`-th tree as:
|
||||
|
||||
.. math::
|
||||
|
||||
\text{obj}^{(t)} &\approx \sum_{i=1}^n [g_i w_{q(x_i)} + \frac{1}{2} h_i w_{q(x_i)}^2] + \gamma T + \frac{1}{2}\lambda \sum_{j=1}^T w_j^2\\
|
||||
&= \sum^T_{j=1} [(\sum_{i\in I_j} g_i) w_j + \frac{1}{2} (\sum_{i\in I_j} h_i + \lambda) w_j^2 ] + \gamma T
|
||||
|
||||
where :math:`I_j = \{i|q(x_i)=j\}` is the set of indices of data points assigned to the :math:`j`-th leaf.
|
||||
Notice that in the second line we have changed the index of the summation because all the data points on the same leaf get the same score.
|
||||
We could further compress the expression by defining :math:`G_j = \sum_{i\in I_j} g_i` and :math:`H_j = \sum_{i\in I_j} h_i`:
|
||||
|
||||
.. math::
|
||||
|
||||
\text{obj}^{(t)} = \sum^T_{j=1} [G_jw_j + \frac{1}{2} (H_j+\lambda) w_j^2] +\gamma T
|
||||
|
||||
In this equation, :math:`w_j` are independent with respect to each other, the form :math:`G_jw_j+\frac{1}{2}(H_j+\lambda)w_j^2` is quadratic and the best :math:`w_j` for a given structure :math:`q(x)` and the best objective reduction we can get is:
|
||||
|
||||
.. math::
|
||||
|
||||
w_j^\ast &= -\frac{G_j}{H_j+\lambda}\\
|
||||
\text{obj}^\ast &= -\frac{1}{2} \sum_{j=1}^T \frac{G_j^2}{H_j+\lambda} + \gamma T
|
||||
|
||||
The last equation measures *how good* a tree structure :math:`$q(x)` is.
|
||||
|
||||
.. image:: https://raw.githubusercontent.com/dmlc/web-data/master/xgboost/model/struct_score.png
|
||||
:width: 100%
|
||||
:alt: illustration of structure score (fitness)
|
||||
|
||||
If all this sounds a bit complicated, let's take a look at the picture, and see how the scores can be calculated.
|
||||
Basically, for a given tree structure, we push the statistics :math:`g_i` and :math:`h_i` to the leaves they belong to,
|
||||
sum the statistics together, and use the formula to calculate how good the tree is.
|
||||
This score is like the impurity measure in a decision tree, except that it also takes the model complexity into account.
|
||||
|
||||
Learn the tree structure
|
||||
========================
|
||||
Now that we have a way to measure how good a tree is, ideally we would enumerate all possible trees and pick the best one.
|
||||
In practice this is intractable, so we will try to optimize one level of the tree at a time.
|
||||
Specifically we try to split a leaf into two leaves, and the score it gains is
|
||||
|
||||
.. math::
|
||||
Gain = \frac{1}{2} \left[\frac{G_L^2}{H_L+\lambda}+\frac{G_R^2}{H_R+\lambda}-\frac{(G_L+G_R)^2}{H_L+H_R+\lambda}\right] - \gamma
|
||||
|
||||
This formula can be decomposed as 1) the score on the new left leaf 2) the score on the new right leaf 3) The score on the original leaf 4) regularization on the additional leaf.
|
||||
We can see an important fact here: if the gain is smaller than :math:`\gamma`, we would do better not to add that branch. This is exactly the **pruning** techniques in tree based
|
||||
models! By using the principles of supervised learning, we can naturally come up with the reason these techniques work :)
|
||||
|
||||
For real valued data, we usually want to search for an optimal split. To efficiently do so, we place all the instances in sorted order, like the following picture.
|
||||
|
||||
.. image:: https://raw.githubusercontent.com/dmlc/web-data/master/xgboost/model/split_find.png
|
||||
:width: 100%
|
||||
:alt: Schematic of choosing the best split
|
||||
|
||||
A left to right scan is sufficient to calculate the structure score of all possible split solutions, and we can find the best split efficiently.
|
||||
|
||||
**********************
|
||||
Final words on XGBoost
|
||||
**********************
|
||||
Now that you understand what boosted trees are, you may ask, where is the introduction for XGBoost?
|
||||
XGBoost is exactly a tool motivated by the formal principle introduced in this tutorial!
|
||||
More importantly, it is developed with both deep consideration in terms of **systems optimization** and **principles in machine learning**.
|
||||
The goal of this library is to push the extreme of the computation limits of machines to provide a **scalable**, **portable** and **accurate** library.
|
||||
Make sure you try it out, and most importantly, contribute your piece of wisdom (code, examples, tutorials) to the community!
|
||||
@@ -1,90 +0,0 @@
|
||||
Monotonic Constraints
|
||||
=====================
|
||||
|
||||
It is often the case in a modeling problem or project that the functional form of an acceptable model is constrained in some way. This may happen due to business considerations, or because of the type of scientific question being investigated. In some cases, where there is a very strong prior belief that the true relationship has some quality, constraints can be used to improve the predictive performance of the model.
|
||||
|
||||
A common type of constraint in this situation is that certain features bear a *monotonic* relationship to the predicted response:
|
||||
|
||||
```math
|
||||
f(x_1, x_2, \ldots, x, \ldots, x_{n-1}, x_n) \leq f(x_1, x_2, \ldots, x', \ldots, x_{n-1}, x_n)
|
||||
```
|
||||
|
||||
whenever ``$ x \leq x' $`` is an *increasing constraint*; or
|
||||
|
||||
```math
|
||||
f(x_1, x_2, \ldots, x, \ldots, x_{n-1}, x_n) \geq f(x_1, x_2, \ldots, x', \ldots, x_{n-1}, x_n)
|
||||
```
|
||||
|
||||
whenever ``$ x \leq x' $`` is a *decreasing constraint*.
|
||||
|
||||
XGBoost has the ability to enforce monotonicity constraints on any features used in a boosted model.
|
||||
|
||||
A Simple Example
|
||||
----------------
|
||||
|
||||
To illustrate, let's create some simulated data with two features and a response according to the following scheme
|
||||
|
||||
```math
|
||||
y = 5 x_1 + \sin(10 \pi x_1) - 5 x_2 - \cos(10 \pi x_2) + N(0, 0.01)
|
||||
|
||||
x_1, x_2 \in [0, 1]
|
||||
```
|
||||
|
||||
The response generally increases with respect to the ``$ x_1 $`` feature, but a sinusoidal variation has been superimposed, resulting in the true effect being non-monotonic. For the ``$ x_2 $`` feature the variation is decreasing with a sinusoidal variation.
|
||||
|
||||
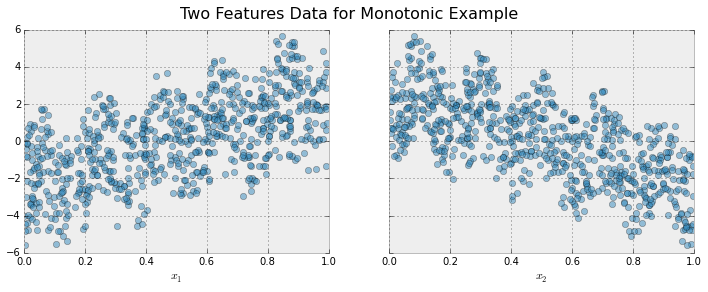
|
||||
|
||||
Let's fit a boosted tree model to this data without imposing any monotonic constraints
|
||||
|
||||
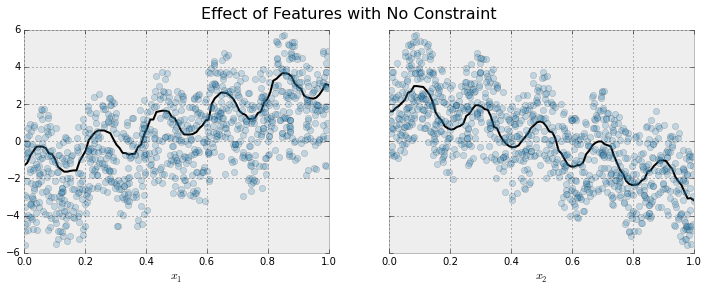
|
||||
|
||||
The black curve shows the trend inferred from the model for each feature. To make these plots the distinguished feature ``$x_i$`` is fed to the model over a one-dimensional grid of values, while all the other features (in this case only one other feature) are set to their average values. We see that the model does a good job of capturing the general trend with the oscillatory wave superimposed.
|
||||
|
||||
Here is the same model, but fit with monotonicity constraints
|
||||
|
||||
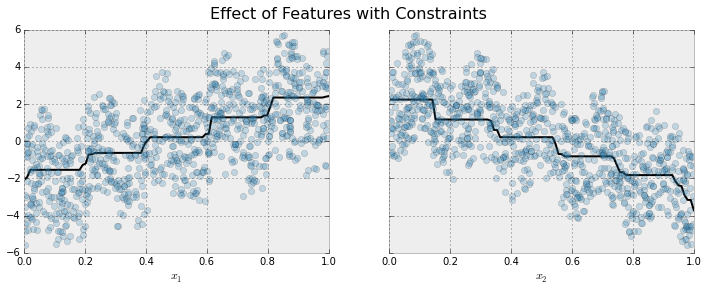
|
||||
|
||||
We see the effect of the constraint. For each variable the general direction of the trend is still evident, but the oscillatory behaviour no longer remains as it would violate our imposed constraints.
|
||||
|
||||
Enforcing Monotonic Constraints in XGBoost
|
||||
------------------------------------------
|
||||
|
||||
It is very simple to enforce monotonicity constraints in XGBoost. Here we will give an example using Python, but the same general idea generalizes to other platforms.
|
||||
|
||||
Suppose the following code fits your model without monotonicity constraints
|
||||
|
||||
```python
|
||||
model_no_constraints = xgb.train(params, dtrain,
|
||||
num_boost_round = 1000, evals = evallist,
|
||||
early_stopping_rounds = 10)
|
||||
```
|
||||
|
||||
Then fitting with monotonicity constraints only requires adding a single parameter
|
||||
|
||||
```python
|
||||
params_constrained = params.copy()
|
||||
params_constrained['monotone_constraints'] = "(1,-1)"
|
||||
|
||||
model_with_constraints = xgb.train(params_constrained, dtrain,
|
||||
num_boost_round = 1000, evals = evallist,
|
||||
early_stopping_rounds = 10)
|
||||
```
|
||||
|
||||
In this example the training data ```X``` has two columns, and by using the parameter values ```(1,-1)``` we are telling XGBoost to impose an increasing constraint on the first predictor and a decreasing constraint on the second.
|
||||
|
||||
Some other examples:
|
||||
|
||||
- ```(1,0)```: An increasing constraint on the first predictor and no constraint on the second.
|
||||
- ```(0,-1)```: No constraint on the first predictor and a decreasing constraint on the second.
|
||||
|
||||
**Choise of tree construction algorithm**. To use monotonic constraints, be
|
||||
sure to set the `tree_method` parameter to one of `'exact'`, `'hist'`, and
|
||||
`'gpu_hist'`.
|
||||
|
||||
**Note for the `'hist'` tree construction algorithm**.
|
||||
If `tree_method` is set to either `'hist'` or `'gpu_hist'`, enabling monotonic
|
||||
constraints may produce unnecessarily shallow trees. This is because the
|
||||
`'hist'` method reduces the number of candidate splits to be considered at each
|
||||
split. Monotonic constraints may wipe out all available split candidates, in
|
||||
which case no split is made. To reduce the effect, you may want to increase
|
||||
the `max_bin` parameter to consider more split candidates.
|
||||
95
doc/tutorials/monotonic.rst
Normal file
95
doc/tutorials/monotonic.rst
Normal file
@@ -0,0 +1,95 @@
|
||||
#####################
|
||||
Monotonic Constraints
|
||||
#####################
|
||||
|
||||
It is often the case in a modeling problem or project that the functional form of an acceptable model is constrained in some way. This may happen due to business considerations, or because of the type of scientific question being investigated. In some cases, where there is a very strong prior belief that the true relationship has some quality, constraints can be used to improve the predictive performance of the model.
|
||||
|
||||
A common type of constraint in this situation is that certain features bear a **monotonic** relationship to the predicted response:
|
||||
|
||||
.. math::
|
||||
|
||||
f(x_1, x_2, \ldots, x, \ldots, x_{n-1}, x_n) \leq f(x_1, x_2, \ldots, x', \ldots, x_{n-1}, x_n)
|
||||
|
||||
whenever :math:`x \leq x'` is an **increasing constraint**; or
|
||||
|
||||
.. math::
|
||||
|
||||
f(x_1, x_2, \ldots, x, \ldots, x_{n-1}, x_n) \geq f(x_1, x_2, \ldots, x', \ldots, x_{n-1}, x_n)
|
||||
|
||||
whenever :math:`x \leq x'` is a **decreasing constraint**.
|
||||
|
||||
XGBoost has the ability to enforce monotonicity constraints on any features used in a boosted model.
|
||||
|
||||
****************
|
||||
A Simple Example
|
||||
****************
|
||||
|
||||
To illustrate, let's create some simulated data with two features and a response according to the following scheme
|
||||
|
||||
.. math::
|
||||
|
||||
y = 5 x_1 + \sin(10 \pi x_1) - 5 x_2 - \cos(10 \pi x_2) + N(0, 0.01)
|
||||
x_1, x_2 \in [0, 1]
|
||||
|
||||
The response generally increases with respect to the :math:`x_1` feature, but a sinusoidal variation has been superimposed, resulting in the true effect being non-monotonic. For the :math:`x_2` feature the variation is decreasing with a sinusoidal variation.
|
||||
|
||||
.. image:: https://raw.githubusercontent.com/dmlc/web-data/master/xgboost/monotonic/two.feature.sample.data.png
|
||||
:alt: Data in sinusoidal fit
|
||||
|
||||
Let's fit a boosted tree model to this data without imposing any monotonic constraints:
|
||||
|
||||
.. image:: https://raw.githubusercontent.com/dmlc/web-data/master/xgboost/monotonic/two.feature.no.constraint.png
|
||||
:alt: Fit of Model with No Constraint
|
||||
|
||||
The black curve shows the trend inferred from the model for each feature. To make these plots the distinguished feature :math:`x_i` is fed to the model over a one-dimensional grid of values, while all the other features (in this case only one other feature) are set to their average values. We see that the model does a good job of capturing the general trend with the oscillatory wave superimposed.
|
||||
|
||||
Here is the same model, but fit with monotonicity constraints:
|
||||
|
||||
.. image:: https://raw.githubusercontent.com/dmlc/web-data/master/xgboost/monotonic/two.feature.with.constraint.png
|
||||
:alt: Fit of Model with Constraint
|
||||
|
||||
We see the effect of the constraint. For each variable the general direction of the trend is still evident, but the oscillatory behaviour no longer remains as it would violate our imposed constraints.
|
||||
|
||||
******************************************
|
||||
Enforcing Monotonic Constraints in XGBoost
|
||||
******************************************
|
||||
|
||||
It is very simple to enforce monotonicity constraints in XGBoost. Here we will give an example using Python, but the same general idea generalizes to other platforms.
|
||||
|
||||
Suppose the following code fits your model without monotonicity constraints
|
||||
|
||||
.. code-block:: python
|
||||
|
||||
model_no_constraints = xgb.train(params, dtrain,
|
||||
num_boost_round = 1000, evals = evallist,
|
||||
early_stopping_rounds = 10)
|
||||
|
||||
Then fitting with monotonicity constraints only requires adding a single parameter
|
||||
|
||||
.. code-block:: python
|
||||
|
||||
params_constrained = params.copy()
|
||||
params_constrained['monotone_constraints'] = "(1,-1)"
|
||||
|
||||
model_with_constraints = xgb.train(params_constrained, dtrain,
|
||||
num_boost_round = 1000, evals = evallist,
|
||||
early_stopping_rounds = 10)
|
||||
|
||||
In this example the training data ``X`` has two columns, and by using the parameter values ``(1,-1)`` we are telling XGBoost to impose an increasing constraint on the first predictor and a decreasing constraint on the second.
|
||||
|
||||
Some other examples:
|
||||
|
||||
- ``(1,0)``: An increasing constraint on the first predictor and no constraint on the second.
|
||||
- ``(0,-1)``: No constraint on the first predictor and a decreasing constraint on the second.
|
||||
|
||||
**Choise of tree construction algorithm**. To use monotonic constraints, be
|
||||
sure to set the ``tree_method`` parameter to one of ``exact``, ``hist``, and
|
||||
``gpu_hist``.
|
||||
|
||||
**Note for the 'hist' tree construction algorithm**.
|
||||
If ``tree_method`` is set to either ``hist`` or ``gpu_hist``, enabling monotonic
|
||||
constraints may produce unnecessarily shallow trees. This is because the
|
||||
``hist`` method reduces the number of candidate splits to be considered at each
|
||||
split. Monotonic constraints may wipe out all available split candidates, in
|
||||
which case no split is made. To reduce the effect, you may want to increase
|
||||
the ``max_bin`` parameter to consider more split candidates.
|
||||
55
doc/tutorials/param_tuning.rst
Normal file
55
doc/tutorials/param_tuning.rst
Normal file
@@ -0,0 +1,55 @@
|
||||
#########################
|
||||
Notes on Parameter Tuning
|
||||
#########################
|
||||
Parameter tuning is a dark art in machine learning, the optimal parameters
|
||||
of a model can depend on many scenarios. So it is impossible to create a
|
||||
comprehensive guide for doing so.
|
||||
|
||||
This document tries to provide some guideline for parameters in XGBoost.
|
||||
|
||||
************************************
|
||||
Understanding Bias-Variance Tradeoff
|
||||
************************************
|
||||
If you take a machine learning or statistics course, this is likely to be one
|
||||
of the most important concepts.
|
||||
When we allow the model to get more complicated (e.g. more depth), the model
|
||||
has better ability to fit the training data, resulting in a less biased model.
|
||||
However, such complicated model requires more data to fit.
|
||||
|
||||
Most of parameters in XGBoost are about bias variance tradeoff. The best model
|
||||
should trade the model complexity with its predictive power carefully.
|
||||
:doc:`Parameters Documentation </parameter>` will tell you whether each parameter
|
||||
ill make the model more conservative or not. This can be used to help you
|
||||
turn the knob between complicated model and simple model.
|
||||
|
||||
*******************
|
||||
Control Overfitting
|
||||
*******************
|
||||
When you observe high training accuracy, but low test accuracy, it is likely that you encountered overfitting problem.
|
||||
|
||||
There are in general two ways that you can control overfitting in XGBoost
|
||||
|
||||
* The first way is to directly control model complexity
|
||||
|
||||
- This include ``max_depth``, ``min_child_weight`` and ``gamma``
|
||||
|
||||
* The second way is to add randomness to make training robust to noise
|
||||
|
||||
- This include ``subsample`` and ``colsample_bytree``.
|
||||
- You can also reduce stepsize ``eta``. Rremember to increase ``num_round`` when you do so.
|
||||
|
||||
*************************
|
||||
Handle Imbalanced Dataset
|
||||
*************************
|
||||
For common cases such as ads clickthrough log, the dataset is extremely imbalanced.
|
||||
This can affect the training of XGBoost model, and there are two ways to improve it.
|
||||
|
||||
* If you care only about the overall performance metric (AUC) of your prediction
|
||||
|
||||
- Balance the positive and negative weights via ``scale_pos_weight``
|
||||
- Use AUC for evaluation
|
||||
|
||||
* If you care about predicting the right probability
|
||||
|
||||
- In such a case, you cannot re-balance the dataset
|
||||
- Set parameter ``max_delta_step`` to a finite number (say 1) to help convergence
|
||||
Reference in New Issue
Block a user